Aluminium »
PDB 7t3b-8ien »
8eew »
Aluminium in PDB 8eew: Cryoem of the Soluble OPA1 Dimer From the Gdp-Alfx Bound Helical Assembly on A Lipid Membrane
Other elements in 8eew:
The structure of Cryoem of the Soluble OPA1 Dimer From the Gdp-Alfx Bound Helical Assembly on A Lipid Membrane also contains other interesting chemical elements:
| Fluorine | (F) | 8 atoms |
| Potassium | (K) | 2 atoms |
| Magnesium | (Mg) | 2 atoms |
Aluminium Binding Sites:
The binding sites of Aluminium atom in the Cryoem of the Soluble OPA1 Dimer From the Gdp-Alfx Bound Helical Assembly on A Lipid Membrane
(pdb code 8eew). This binding sites where shown within
5.0 Angstroms radius around Aluminium atom.
In total 2 binding sites of Aluminium where determined in the Cryoem of the Soluble OPA1 Dimer From the Gdp-Alfx Bound Helical Assembly on A Lipid Membrane, PDB code: 8eew:
Jump to Aluminium binding site number: 1; 2;
In total 2 binding sites of Aluminium where determined in the Cryoem of the Soluble OPA1 Dimer From the Gdp-Alfx Bound Helical Assembly on A Lipid Membrane, PDB code: 8eew:
Jump to Aluminium binding site number: 1; 2;
Aluminium binding site 1 out of 2 in 8eew
Go back to
Aluminium binding site 1 out
of 2 in the Cryoem of the Soluble OPA1 Dimer From the Gdp-Alfx Bound Helical Assembly on A Lipid Membrane
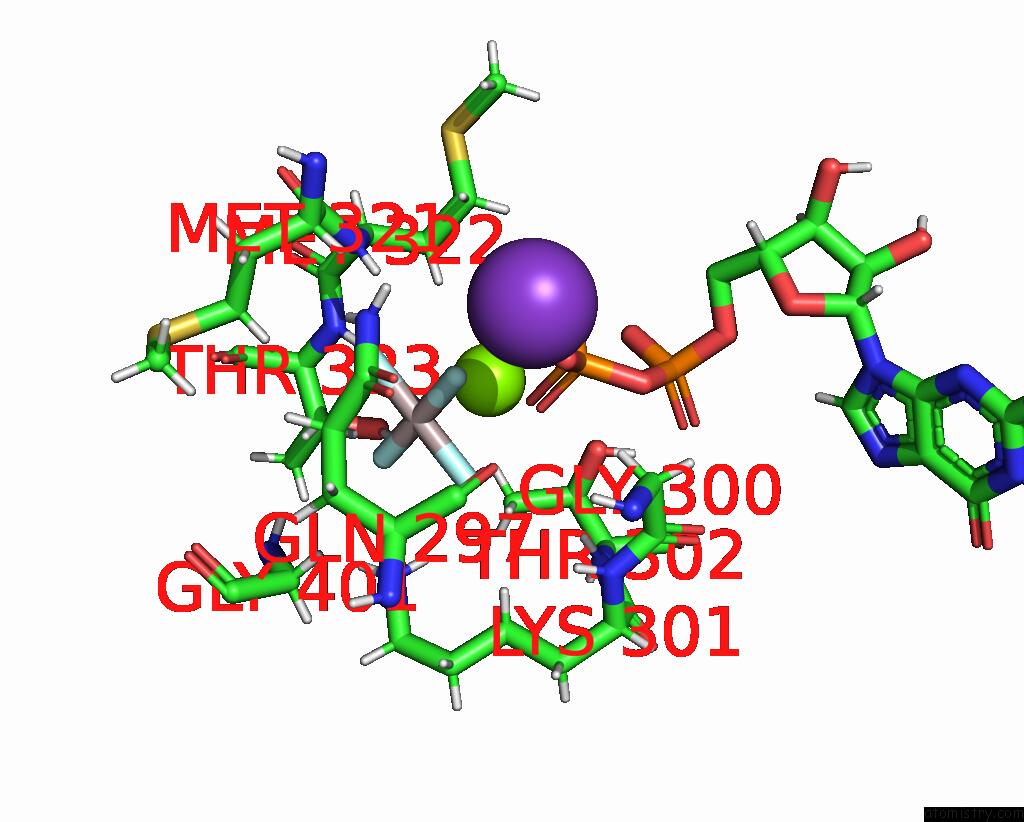
Mono view
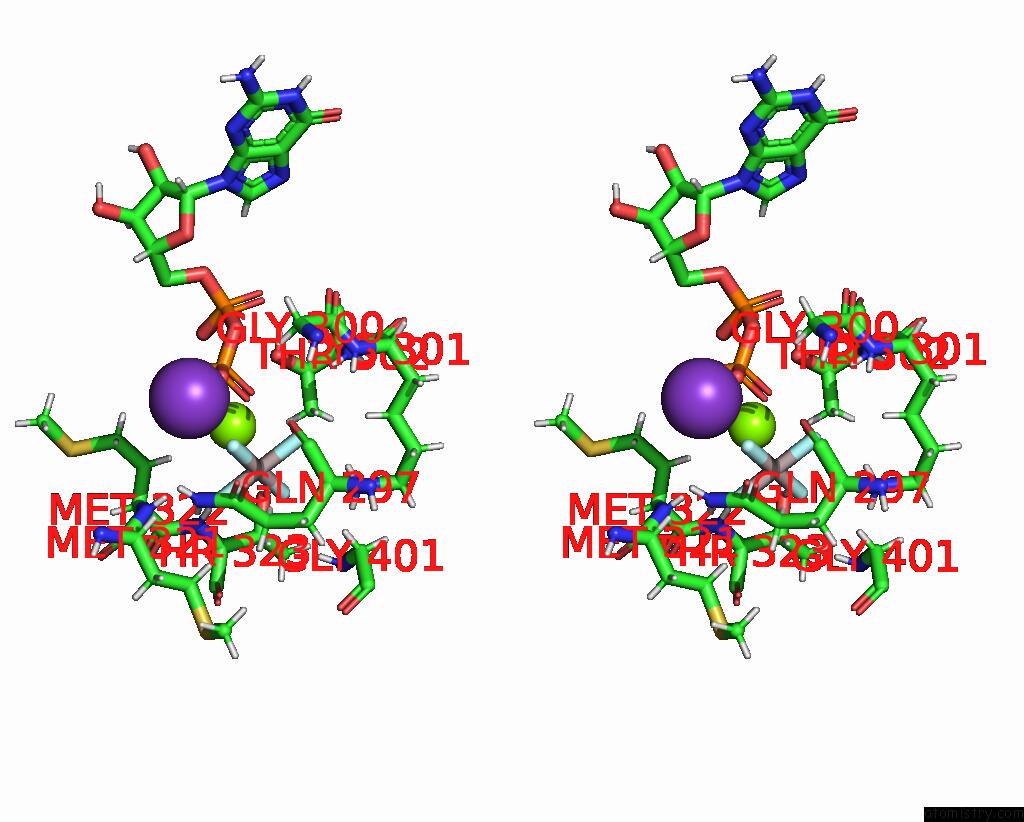
Stereo pair view

Mono view

Stereo pair view
A full contact list of Aluminium with other atoms in the Al binding
site number 1 of Cryoem of the Soluble OPA1 Dimer From the Gdp-Alfx Bound Helical Assembly on A Lipid Membrane within 5.0Å range:
|
Aluminium binding site 2 out of 2 in 8eew
Go back to
Aluminium binding site 2 out
of 2 in the Cryoem of the Soluble OPA1 Dimer From the Gdp-Alfx Bound Helical Assembly on A Lipid Membrane
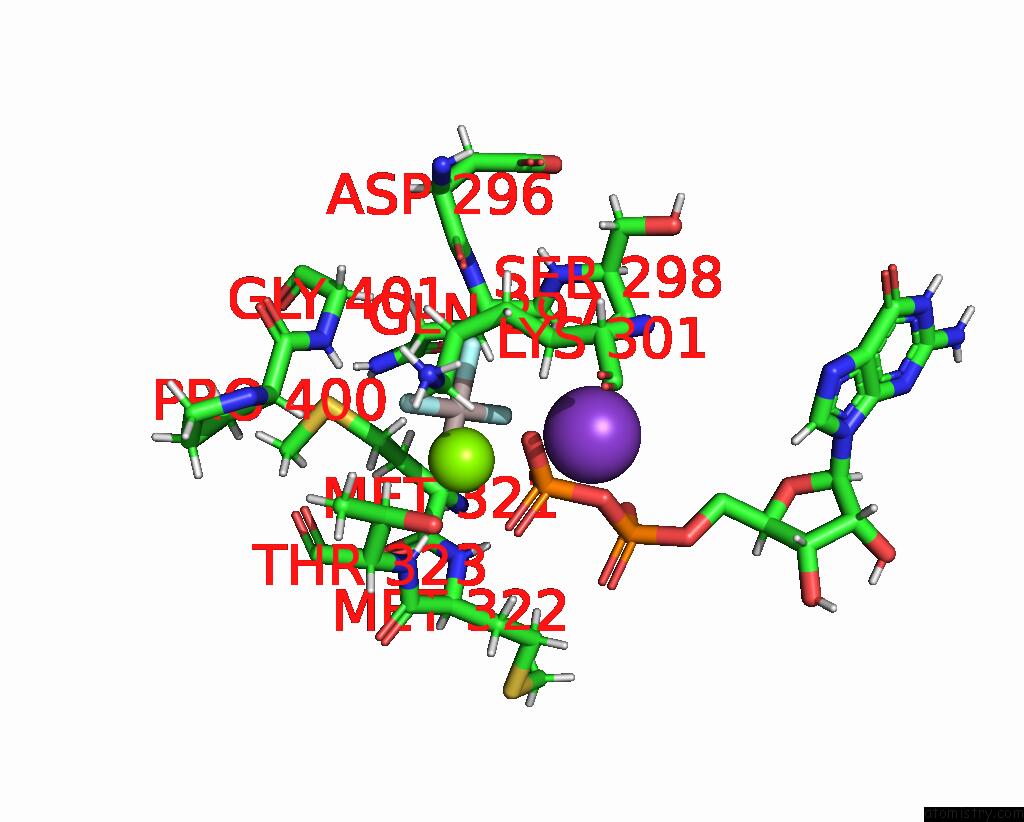
Mono view
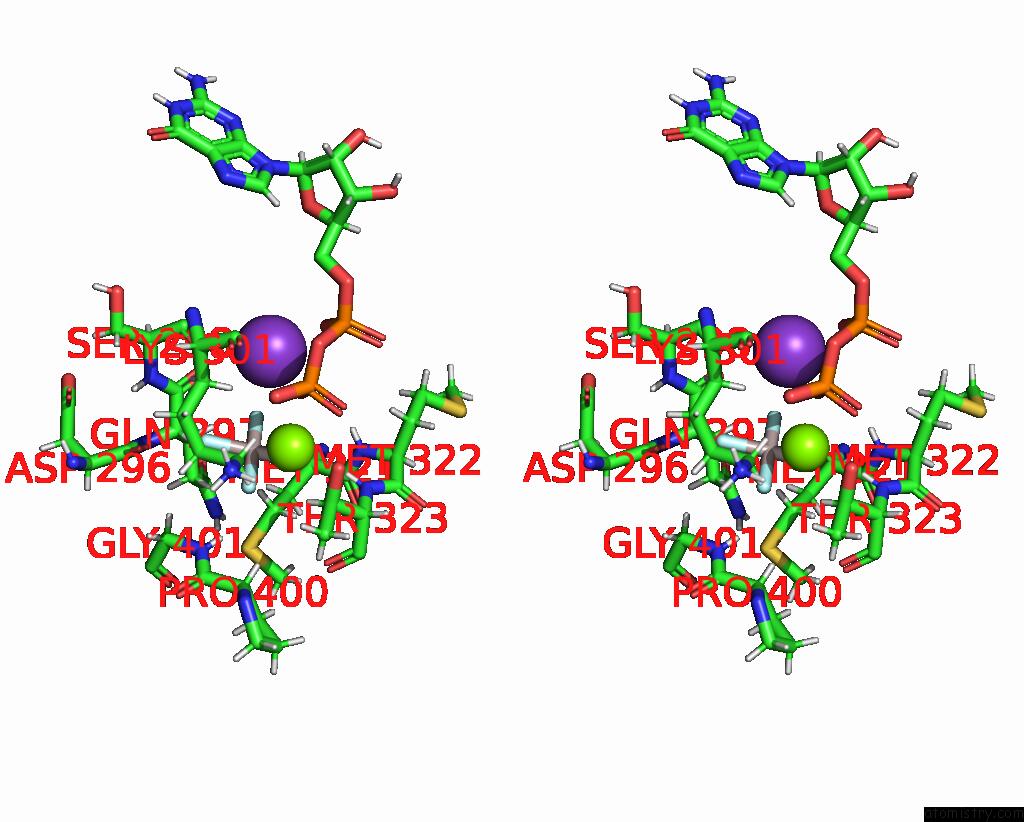
Stereo pair view

Mono view

Stereo pair view
A full contact list of Aluminium with other atoms in the Al binding
site number 2 of Cryoem of the Soluble OPA1 Dimer From the Gdp-Alfx Bound Helical Assembly on A Lipid Membrane within 5.0Å range:
|
Reference:
S.B.Nyenhuis,
X.Wu,
A.E.Stanton,
M.P.Strub,
Y.I.Yim,
B.Canagarajah,
J.E.Hinshaw.
OPA1 Helical Structures Give Perspective to Mitochondrial Dysfunction Nature 2023.
ISSN: ESSN 1476-4687
Page generated: Wed Jul 10 10:17:22 2024
ISSN: ESSN 1476-4687
Last articles
Zn in 9MJ5Zn in 9HNW
Zn in 9G0L
Zn in 9FNE
Zn in 9DZN
Zn in 9E0I
Zn in 9D32
Zn in 9DAK
Zn in 8ZXC
Zn in 8ZUF